Research summary
My current research focus on the mechanisms of epigenetic regulation with a focus on DNA methylation and its regulation by environmental cues. The research is mainly done in a model plant Arabidopsis thaliana.
Regulation of DNA methylation by UV-B light.
Under sunshine with UVB, the UVB photoreceptor UVR8 perceives UVB and undergoes monomerization followed by nuclear import. In the nucleus, UVR8 interacts with DRM2 and induces DNA hypomethylation by inhibiting the catalytic activity and chromatin association of the de novo DNA methyltransferase DRM2. This study has been published in Nature Plants (2021). Following this, we are also studying natural Arabidopsis thaliana and other alpine species to reveal the role of UVB and DNA methylation during plant adaption to the harsh high altitudes.
Under sunshine with UVB, the UVB photoreceptor UVR8 perceives UVB and undergoes monomerization followed by nuclear import. In the nucleus, UVR8 interacts with DRM2 and induces DNA hypomethylation by inhibiting the catalytic activity and chromatin association of the de novo DNA methyltransferase DRM2. This study has been published in Nature Plants (2021). Following this, we are also studying natural Arabidopsis thaliana and other alpine species to reveal the role of UVB and DNA methylation during plant adaption to the harsh high altitudes.
Mechanism and evolution of non-CG DNA methylation in plants.
(1) Mechanism of DNA methylation catalyzed by the DRM2 in RNA-directed DNA methylation (RdDM) pathway.
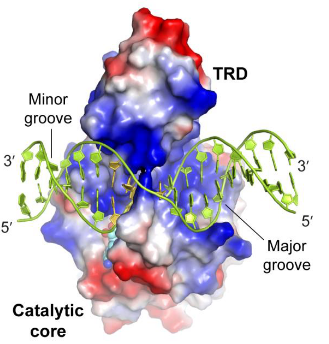
By collaborating with Dr. Jikui Song at University of California-Riverside, we revealed the structure-function characterizations of DRM2-mediated methylation. An arginine finger from the catalytic loop intercalates into DNA minor groove, inducing large DNA deformation that impacts the substrate specificity of DRM2. Mutating DRM2 could result shift the substrate preference from CHH to CHG and CG (H=A,T,C).
(2) Mechanism of H3K9me2 dependent non-CG DNA methylation by CMT methyltransferases.
We are revealing the mechanism of CMT2 and CMT3 mediated non-CG DNA methylation by using a combination of structural, biochemical, and genetic approaches. This type of DNA methylation plays key roles in silencing transposons and safeguarding the genome, and has been conserved in plant for more than 800 million years on earth.
(1) Mechanism of DNA methylation catalyzed by the DRM2 in RNA-directed DNA methylation (RdDM) pathway.
By collaborating with Dr. Jikui Song at University of California-Riverside, we revealed the structure-function characterizations of DRM2-mediated methylation. An arginine finger from the catalytic loop intercalates into DNA minor groove, inducing large DNA deformation that impacts the substrate specificity of DRM2. Mutating DRM2 could result shift the substrate preference from CHH to CHG and CG (H=A,T,C).
(2) Mechanism of H3K9me2 dependent non-CG DNA methylation by CMT methyltransferases.
We are revealing the mechanism of CMT2 and CMT3 mediated non-CG DNA methylation by using a combination of structural, biochemical, and genetic approaches. This type of DNA methylation plays key roles in silencing transposons and safeguarding the genome, and has been conserved in plant for more than 800 million years on earth.